IDENTIFICATION AND PREDICTION OF THE COLOR COAT IN THE EQUINE SPECIE
|
Description |
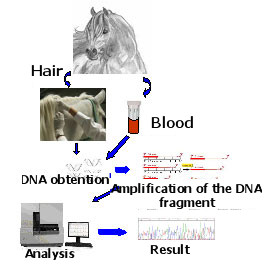
Scheme of the methodology used for genetic identification.
The identification and prediction of the coat color in horses is based on the identification of the gene or genes that are involved and control the hair color and skin.
From blood or hair samples the DNA is obtained and we proceed to identify genotypes for each gene, which will reveal the Genetic Composition of an animal or group of animals and predict the color of the offspring that would be obtained from certain crossings.
At this time the genes involved in basic colorations (black, sorrel or chestnut), the gene responsible for some of the dilutions of interest (bay, palomino, black ashen, pearl, cremello, cream ashen and silver) and those responsible for the appearance of some pints (tobiano, sabino, overo) are known.
|
How does it work |
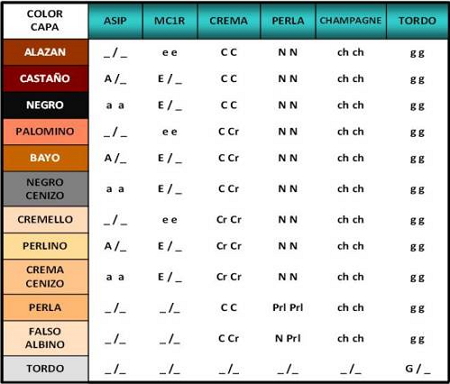
Table1. Genotypes combinations for the principal layer colorations in equine species.
For the analysis of the identification of the genes that are responsible of the color coat; simple, fast and inexpensive techniques can be performed from any sample containing nucleated cells, for example, blood or hair bulb. The procedure is as follows: first we carry out a DNA isolation and next we perform an enzymatic amplification of the gene by the PCR technique to identify mutations, deletions and/or insertions that have been inherited from one or both parents.
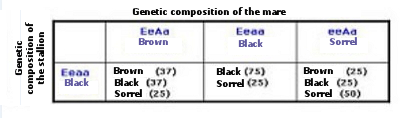
Depending on the gene we want to analyze, different techniques are used, such as sequencing, SSCP or real-time PCR. When the genotypes that are acting on the basic coat (black, chestnut, sorrel), diluted (bay, palomino, black ashen, pearl, pearl, cremello, cream ashen, silver) and pints (tobiano, sabino, overo) are identified, it will be possible to know the color coat that an animal will transmit to their offspring. Table 1 shows the probability of getting offspring with a chestnut, black or sorrel coat depending on the genetic composition of the parents for the Extension (Ee) and Agouti (Aa) genes.
For example, if a black coat mare with genotype Eeaa, and a stallion also with a black coat and with the same genotype are intersected, the offspring will have black cloak with a probability of 75%, but the 25% offspring will have a sorrel coat.
|
Advantages |
The identification of genes and genotypes that the horses carry allows designing matings in order to achieve an offspring with the desired coat, even when the base coat can be masked by the action of a dominant gene as the gray coat (locus gray) which is dominant over the rest of the colorations and do not allow to know what the coat that can transmit to their offspring are. The ease, quickness of determination and low cost make it ideal to be applied widely by breeders' associations or individual breeders.
|
Where has it been developed |
This technology has been developed by the Genetic Service at the Faculty of Veterinary Medicine, Universidad Complutense de Madrid, by a research group with an extensive participation in national and european projects of research that have allowed them to develop the methodologies necessary for this offer. This center has been offering their services since 1996.
Since its launch, the service of genetic identification color coat is being used by many Purebred Spanish horse breeders.
The technological basis of this Genetic Service is supported by the research that the authors have conducted over the past 20 years and spread through numerous scientific publications in relevant journals, counting also with extensive experience in collaborations with companies and associations. They are part of the group "Animal Nutrigenomics" of the Animal Production Department, Faculty of Veterinary Medicine of the UCM, directed by Susana Dunner. This research group is included in the cluster of Agri-food and Health of the International Excellence Campus (CEI-Moncloa).
|
And also |
The Genetic Service of the Veterinary Medicine Faculty of Madrid, offers support services to daily clinical activities of veterinary (parental controls, genetic identification, sexing birds by molecular techniques, diagnosis of carriers of hereditary diseases,…) and other professionals for other purposes (genetic evaluation, estimation of genetic parameters, etc.).
The knowledge that we are having about the genomes of species of domestic animals, allows us to identify a number of genes that can be used in many applications of interest, such as the choice of breeding animals with free responsible genes.
|
Contact |
|
© Office for the Transfer of Research Results – UCM |
|
PDF Downloads |
|
Classification |
|
Responsible Researcher |
Javier Cañón Ferreras: genetica@vet.ucm.es
Department: Animal Production
Faculty: Veterinary Medicine